-
PDF
- Split View
-
Views
-
Cite
Cite
Triinu Koressaar, Maido Remm, Enhancements and modifications of primer design program Primer3, Bioinformatics, Volume 23, Issue 10, May 2007, Pages 1289–1291, https://doi.org/10.1093/bioinformatics/btm091
Close - Share Icon Share
Abstract
Summary: The determination of annealing temperature is a critical step in PCR design. This parameter is typically derived from the melting temperature of the PCR primers, so for successful PCR work it is important to determine the melting temperature of primer accurately. We introduced several enhancements in the widely used primer design program Primer3. The improvements include a formula for calculating melting temperature and a salt correction formula. Also, the new version can take into account the effects of divalent cations, which are included in most PCR buffers. Another modification enables using lowercase masked template sequences for primer design.
Availability: Features described in this article have been implemented into the development code of Primer3 and will be available in future versions (version 1.1 and newer) of Primer3. Also, a modified version is compiled under the name of mPrimer3 which is distributed independently. The web-based version of mPrimer3 is available at http://bioinfo.ebc.ee/mprimer3/ and the binary code is freely downloadable from the URL http://bioinfo.ebc.ee/download/.
Contact: maido.remm@ut.ee
1 INTRODUCTION
Primer3 (Rozen et al., 2000) has been widely used for PCR primer design for more than a decade. The original version uses a table of thermodynamic parameters published by the Breslauer group (Breslauer et al., 1986) and a formula for melting temperature (Tm) calculations published by Rychlik and co-workers (Rychlik et al., 1990). The outdated table of thermodynamic parameters and method for calculating Tm are considered the main drawback of Primer3 (Chavali et al., 2005). Improved sets of thermodynamic parameters and improved formulae are now available (Owczarzy et al., 1998; Panjkovich et al., 2005; SantaLucia, 1998; SantaLucia et al., 2004). An important influence on primer melting temperature is the concentration of mono- and divalent cations in the solution. As nearest-neighbor parameters are measured in a specific salt concentration (e.g. 1 M NaCl), the melting temperature should be corrected for the actual conditions of the PCR buffer. The original Primer3 release uses a salt correction formula published by Schildkraut and Lifson (Schildkraut and Lifson, 1965), which does not consider the effect of divalent cations on melting temperature calculations. Many other widely used general primer design programs like OLIGO6 (http://www.oligo.net) and PRIDE (Haas et al., 1998) similarly use outdated formulas for Tm calculation. Many other primer design software exist which already use up-to-date table of thermodynamic parameters for melting temperature calculation, e.g. PrimerPremier (http://www.premierbiosoft.com/), FastPCR (http://www.biocenter.helsinki.fi/bi/Programs/fastpcr.htm) or ORFprimer (http://www.proteinstrukturfabrik.de/ORFprimer). However, from the set of programs with up-to-date thermodynamic parameters only few of them (FastPCR, ORFprimer) allow performing automatic command-line calculations for large datasets similarly to Primer3. Another problem with the current version Primer3 appears if one needs to design PCR primers to masked sequences. Primer3 (release 1.0) is optimized for use with the masking program RepeatMasker (Smith A.F.A, http://www.repeatmasker.org/), which masks the sequence with a series of ‘N’ characters in repeat regions if used with default parameters. By default, Primer3 excludes any primer candidates that contain ‘N’ characters, which is a reasonable way of avoiding most common repeats. However, other masking methods may mask only short stretches of the template DNA sequence. For example, DUST (Morgulis et al., 2006), Windowmasker (Morgulis et al., 2006) and GenomeMasker (Andreson et al., 2006a) mask much shorter regions and occasionally only one nucleotide. Furthermore, some of these short masked features are deleterious only if they happen to overlap the 3′ end of the primer and can be tolerated when they overlap the 5′ end. If the target sequence is masked with ‘N’ characters, the design of primers comprising ‘N’ characters is typically forbidden. As far as we know, none of the existing primer design programs supports the usage of soft-masked (lowercase-masked) target sequences. Therefore, we have implemented a new feature into mPrimer3 that allows primers to be designed from lowercase masked sequences. Lowercase masking preserves the DNA sequence and allows primers to be designed that partly overlap the masked region.
2 MODIFICATIONS OF THE ALGORITHM
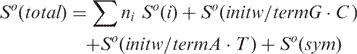
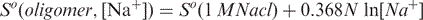
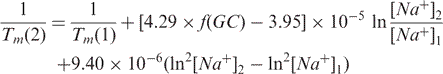
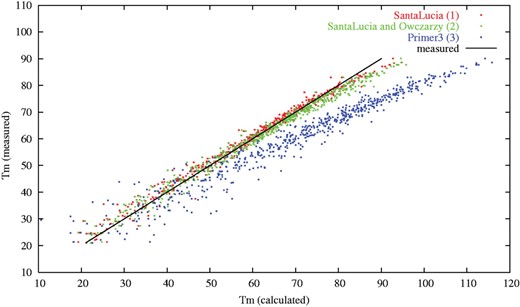
The correlation between predicted and measured melting temperatures for three different Tm calculation methods. Methods (1) and (2) were implemented into the modified version of Primer3. Method (3) corresponds to the original version of Primer3. For primers of typical length (15–30 nucleotides) the average differences between the experimental and predicted Tm were 1.37, 1.78 and 11.70°C for methods (1), (2) and (3) respectively. The experimental melting temperature data used in this analysis were retrieved from the literature (Owczarzy et al., 2004) and include 590 different measurements with 146 different oligonucleotides.
As explained in the introduction, there is sometimes a need to design primers overlapping the masked regions. We have added a new feature to Primer3 allowing primers overlapping lowercase-masked regions to be designed. A novel feature of the modified Primer3 is that primers with a lowercase nucleotide at the 3′ end can be rejected. This behavior relies on the assumption that masked features (e.g. repeats) can partly overlap the primer, but they cannot overlap its 3′ end. Lowercase letters in other positions are accepted, assuming that the masked features do not influence primer performance if they do not overlap the 3′ end.
ACKNOWLEDGEMENTS
This work was partly supported by grant EU19730 from Enterprise Estonia. The authors are grateful to Eric Reppo for initial software changes and Aare Abroi, Priit Palta and Reidar Andreson for critical reading of the manuscript. The authors thank Steve Rozen for advice and friendly support during software development. Funding to pay the Open Access publication charges was provided by the Estonian Ministry of Education and Research grant no. 0182649s04
Conflict of Interest: none declared.
REFERENCES
Author notes
Associate Editor: Limsoon Wong