Abstract
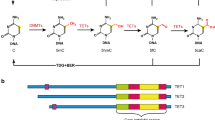
Ten-Eleven Translocation-2 (TET2) inactivation through loss-of-function mutation, deletion and IDH1/2 (Isocitrate Dehydrogenase 1 and 2) gene mutation is a common event in myeloid and lymphoid malignancies. TET2 gene mutations similar to those observed in myeloid and lymphoid malignancies also accumulate with age in otherwise healthy subjects with clonal hematopoiesis. TET2 is one of the three proteins of the TET (Ten-Eleven Translocation) family, which are evolutionarily conserved dioxygenases that catalyze the conversion of 5-methyl-cytosine (5-mC) to 5-hydroxymethyl-cytosine (5-hmC) and promote DNA demethylation. TET dioxygenases require 2-oxoglutarate, oxygen and Fe(II) for their activity, which is enhanced in the presence of ascorbic acid. TET2 is the most expressed TET gene in the hematopoietic tissue, especially in hematopoietic stem cells. In addition to their hydroxylase activity, TET proteins recruit the O-linked β-D-N-acetylglucosamine (O-GlcNAc) transferase (OGT) enzyme to chromatin, which promotes post-transcriptional modifications of histones and facilitates gene expression. The TET2 level is regulated by interaction with IDAX, originating from TET2 gene fission during evolution, and by the microRNA miR-22. TET2 has pleiotropic roles during hematopoiesis, including stem-cell self-renewal, lineage commitment and terminal differentiation of monocytes. Analysis of Tet2 knockout mice, which are viable and fertile, demonstrated that Tet2 functions as a tumor suppressor whose haploinsufficiency initiates myeloid and lymphoid transformations. This review summarizes the recently identified TET2 physiological and pathological functions and discusses how this knowledge influences our therapeutic approaches in hematological malignancies and possibly other tumor types.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Massé A et al. Mutation in TET2 in myeloid cancers. N Engl J Med 2009; 360: 2289–2301.
Langemeijer SM, Kuiper RP, Berends M, Knops R, Aslanyan MG, Massop M et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet 2009; 41: 838–842.
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009; 324: 930–935.
Ono R, Taki T, Taketani T, Taniwaki M, Kobayashi H, Hayashi Y . LCX, leukemia-associated protein with a CXXC domain, is fused to MLL in acute myeloid leukemia with trilineage dysplasia having t(10;11)(q22;q23). Cancer Res 2002; 62: 4075–4080.
Lorsbach RB, Moore J, Mathew S, Raimondi SC, Mukatira ST, Downing JR . TET1, a member of a novel protein family, is fused to MLL in acute myeloid leukemia containing the t(10;11)(q22;q23). Leukemia 2003; 17: 637–641.
Ko M, An J, Bandukwala HS, Chavez L, Aijö T, Pastor WA et al. Modulation of TET2 expression and 5-methylcytosine oxidation by the CXXC domain protein IDAX. Nature 2013; 497: 122–126.
Xu Y, Xu C, Kato A, Tempel W, Abreu JG, Bian C et al. Tet3 CXXC domain and dioxygenase activity cooperatively regulate key genes for Xenopus eye and neural development. Cell 2012; 151: 1200–1213.
Kaelin WG Jr, McKnight SL . Influence of metabolism on epigenetics and disease. Cell 2013; 153: 56–69.
Yin R, Mao SQ, Zhao B, Chong Z, Yang Y, Zhao C et al. Ascorbic Acid enhances tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J Am Chem Soc 2013; 135: 10396–10403.
Blaschke K, Ebata KT, Karimi MM, Zepeda-Martínez JA, Goyal P, Mahapatra S et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 2013; 500: 222–226.
Kovaleva EG, Lipscomb JD . Versatility of biological non-heme Fe(II) centers in oxygen activation reactions. Nat Chem Biol 2008; 4: 186–193.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 2010; 18: 553–567.
Coulter JB, O'Driscoll CM, Bressler JP . Hydroquinone increases 5-hydroxymethylcytosine formation through Ten Eleven Translocation 1 (Tet1) 5-methylcytosine dioxygenase. J Biol Chem 2013; 288: 28792–28800.
Ito S, D’Alessio AC, Taranova OV, Hong K, Sowers LC, Zhang Y . Role of Tet proteins in 5mC to 5-hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 2010; 466: 1129–1133.
Ko M, Huang Y, Jankowska AM, Pape UJ, Tahiliani M, Bandukwala HS et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 2010; 468: 839–843.
Guo JU, Su Y, Zhong C, Ming GL, Song H . Emerging roles of TET proteins and 5-hydroxymethylcytosines in active DNA demethylation and beyond. Cell Cycle 2011; 10: 2662–2668.
Bhutani N, Burns DM, Blau HM . DNA demethylation dynamics. Cell 2011; 146: 866–872.
Nabel CS, Manning SA, Kohli RM . The curious chemical biology of cytosine: deamination, methylation, and oxidation as modulators of genomic potential. ACS Chem Biol 2012; 7: 20–30.
Ito S, Shen L, Dai Q, Wu SC, Collins LB, Swenberg JA et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 2011; 333: 1300–1303.
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011; 333: 1303–1307.
Maiti A, Drohat AC, Thymine DNA . glycosylase can rapidly excise 5-formylcytosine and 5 carboxylcytosine: potential implications for active demethylation of CpG sites. J Biol Chem 2011; 286: 35334–35338.
Cortellino S, Xu J, Sannai M, Moore R, Caretti E, Cigliano A et al. Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 2011; 146: 67–79.
Shen L, Wu H, Diep D, Yamaguchi S, D'Alessio AC, Fung HL et al. Genome-wide analysis reveals TET- and TDG-dependent 5-methylcytosine oxidation dynamics. Cell 2013; 153: 692–706.
Frauer C, Hoffmann T, Bultmann S, Casa V, Cardoso MC, Antes I et al. Recognition of 5-hydroxymethylcytosine by the Uhrf1 SRA domain. PLoS One 2011; 6: e21306.
Yildirim O, Li R, Hung JH, Chen PB, Dong X, Ee LS et al. Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 2011; 147: 1498–1510.
Mellén M, Ayata P, Dewell S, Kriaucionis S, Heintz N . MeCP2 binds to 5-hmC enriched within active genes and accessible chromatin in the nervous system. Cell 2012; 151: 1417–1430.
Spruijt CG, Gnerlich F, Smits AH, Pfaffeneder T, Jansen PW, Bauer C et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 2013; 152: 1146–1159.
Hanover JA, Krause MW, Love DC . Bittersweet memories: linking metabolism to epigenetics through O-GlcNAcylation. Nat Rev Mol Cell Biol 2012; 13: 312–321.
Deplus R, Delatte B, Schwinn MK, Defrance M, Méndez J, Murphy N et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J 2013; 32: 645–655.
Chen Q, Chen Y, Bian C, Fujiki R, Yu X . TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 2013; 493: 561–564.
Vella P, Scelfo A, Jammula S, Chiacchiera F, Williams K, Cuomo A et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol Cell 2013; 49: 645–656.
Pendino F, Nguyen E, Jonassen I, Dysvik B, Azouz A, Lanotte M et al. Functional involvement of RINF, retinoid-inducible nuclear factor (CXXC5), in normal and tumoral human myelopoiesis. Blood 2009; 113: 3172–3181.
Kojima T, Shimazui T, Hinotsu S, Joraku A, Oikawa T, Kawai K et al. Decreased expression of CXXC4 promotes a malignant phenotype in renal cell carcinoma by activating Wnt signaling. Oncogene 2009; 28: 297–305.
Hino S, Kishida S, Michiue T, Fukui A, Sakamoto I, Takada S et al. Inhibition of the Wnt signaling pathway by Idax, a novel Dvl-binding protein. Mol Cell Biol 2001; 21: 330–342.
Guilhamon P, Eskandarpour M, Halai D, Wilson GA, Feber A, Teschendorff AE et al. Meta-analysis of IDH-mutant cancers identifies EBF1 as an interaction partner for TET2. Nat Commun 2013; 4: 2166.
Arioka Y, Watanabe A, Saito K, Yamada Y . Activation-induced cytidine deaminase alters the subcellular localization of Tet family proteins. PLoS One 2012; 7: e45031.
Song SJ, Ito K, Ala U, Kats L, Webster K, Sun SM et al. The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell Stem Cell 2013; 13: 87–101.
Ficz G, Branco MR, Seisenberger S, Santos F, Krueger F, Hore TA et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 2011; 473: 398–402.
Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 2011; 8: 200–213.
Doege CA, Inoue K, Yamashita T, Rhee DB, Travis S, Fujita R et al. Early-stage epigenetic modification during somatic cell reprogramming by Parp1 and Tet2. Nature 2012; 488: 652–655.
Piccolo FM, Bagci H, Brown KE, Landeira D, Soza-Ried J, Feytout A et al. Different roles for Tet1 and Tet2 proteins in reprogramming-mediated erasure of imprints induced by EGC fusion. Mol Cell 2013; 49: 1023–1033.
Xu Y, Wu F, Tan L, Kong L, Xiong L, Deng J et al. Genome-wide regulation of 5-hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell 2011; 42: 451–464.
Williams K, Christensen J, Pedersen MT, Johansen JV, Cloos PA, Rappsilber J et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 2011; 473: 343–348.
Pastor WA, Pape UJ, Huang Y, Henderson HR, Lister R, Ko M et al. Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 2011; 473: 394–397.
Wu H, D'Alessio AC, Ito S, Wang Z, Cui K, Zhao K et al. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev 2011; 25: 679–684.
Wu H, Zhang Y . Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev 2011; 25: 2436–2452.
Williams K, Christensen J, Helin K . DNA methylation: TET proteins-guardians of CpG islands? EMBO Rep 2011; 13: 28–35.
Wu Y, Guo Z, Liu Y, Tang B, Wang Y, Yang L et al. Oct4 and the small molecule inhibitor, SC1, regulates Tet2 expression in mouse embryonic stem cells. Mol Biol Rep 2013; 40: 2897–2906.
Costa Y, Ding J, Theunissen TW, Faiola F, Hore TA, Shliaha PV et al. NANOG-dependent function of TET1 and TET2 in establishment of pluripotency. Nature 2013; 495: 370–374.
Wang T, Wu H, Li Y, Szulwach KE, Lin L, Li X et al. Subtelomeric hotspots of aberrant 5-hydroxymethylcytosine-mediated epigenetic modifications during reprogramming to pluripotency. Nat Cell Biol 2013; 15: 700–711.
Dawlaty MM, Ganz K, Powell BE, Hu YC, Markoulaki S, Cheng AW et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 2011; 9: 166–175.
Dawlaty MM, Breiling A, Le T, Raddatz G, Barrasa MI, Cheng AW et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev Cell 2013; 24: 310–323.
Figueroa ME, Lugthart S, Li Y, Erpelinck-Verschueren C, Deng X, Christos PJ et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell 2010; 17: 13–27.
Moran-Crusio K, Reavie L, Shih A, Abdel-Wahab O, Ndiaye-Lobry D, Lobry C et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 2011; 20: 11–24.
Bocker MT, Tuorto F, Raddatz G, Musch T, Yang FC, Xu M et al. Hydroxylation of 5-methylcytosine by TET2 maintains the active state of the mammalian HOXA cluster. Nat Commun 2012; 3: 818.
Ko M, Bandukwala HS, An J, Lamperti ED, Thompson EC, Hastie R et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc Natl Acad Sci USA 2011; 108: 14566–14571.
Pronier E, Almire C, Mokrani H, Vasanthakumar A, Simon A, da Costa Reis Monte Mor B et al. Inhibition of TET2-mediated conversion of 5-methylcytosine to 5-hydroxymethylcytosine disturbs erythroid and granulomonocytic differentiation of human hematopoietic progenitors. Blood 2011; 118: 2551–2555.
Itzykson R, Kosmider O, Renneville A, Gelsi-Boyer V, Meggendorfer M, Morabito M et al. Prognostic score including gene mutations in chronic myelomonocytic leukemia. J Clin Oncol 2013; 31: 2428–2436.
Itzykson R, Kosmider O, Renneville A, Morabito M, Preudhomme C, Berthon C et al. Clonal architecture of chronic myelomonocytic leukemias. Blood 2013; 121: 2186–2198.
Itzykson R, Solary E . An evolutionary perspective on chronic myelomonocytic leukemia. Leukemia 2013; 27: 1441–1450.
Kallin EM, Rodríguez-Ubreva J, Christensen J, Cimmino L, Aifantis I, Helin K et al. Tet2 facilitates the derepression of myeloid target genes during CEBPα-induced transdifferentiation of pre-B cells. Mol Cell 2012; 48: 266–276.
Klug M, Schmidhofer S, Gebhard C, Andreesen R, Rehli M . 5-Hydroxymethylcytosine is an essential intermediate of active DNA demethylation processes in primary human monocytes. Genome Biol 2013; 14: R46.
Li Z, Cai X, Cai CL, Wang J, Zhang W, Petersen BE et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 2011; 118: 4509–4518.
Quivoron C, Couronné L, Della Valle V, Lopez CK, Plo I, Wagner-Ballon O et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell 2011; 20: 25–38.
Lobry C, Ntziachristos P, Ndiaye-Lobry D, Oh P, Cimmino L, Zhu N et al. Notch pathway activation targets AML-initiating cell homeostasis and differentiation. J Exp Med 2013; 210: 301–319.
Aucagne R, Droin N, Paggetti J, Lagrange B, Largeot A, Hammann A et al. Transcription intermediary factor 1γ is a tumor suppressor in mouse and human chronic myelomonocytic leukemia. J Clin Invest 2011; 121: 2361–2370.
Ward PS, Patel J, Wise DR, Abdel-Wahab O, Bennett BD, Coller HA et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 2010; 17: 225–234.
Koivunen P, Lee S, Duncan CG, Lopez G, Lu G, Ramkissoon S et al. Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature 2012; 483: 484–488.
Lian CG, Xu Y, Ceol C, Wu F, Larson A, Dresser K et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 2012; 50: 1135–1146.
Gambichler T, Sand M, Skrygan M . Loss of 5-hydroxymethylcytosine and ten-eleven translocation 2 protein expression in malignant melanoma. Melanoma Res 2013; 23: 218–220.
Busque L, Patel JP, Figueroa ME, Vasanthakumar A, Provost S, Hamilou Z et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat Genet 2012; 44: 1179–1181.
Sakaguchi H, Okuno Y, Muramatsu H, Yoshida K, Shiraishi Y, Takahashi M et al. Exome sequencing identifies secondary mutations of SETBP1 and JAK3 in juvenile myelomonocytic leukemia. Nat Genet 2013; 45: 937–941.
Langemeijer SM, Jansen JH, Hooijer J, van Hoogen P, Stevens-Linders E, Massop M et al. TET2 mutations in childhood leukemia. Leukemia 2011; 25: 189–192.
Metzeler KH, Maharry K, Radmacher MD, Mrózek K, Margeson D, Becker H et al. TET2 mutations improve the new European LeukemiaNet risk classification of acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol 2011; 29: 1373–1381.
Chan SM, Majeti R . Role of DNMT3A, TET2, and IDH1/2 mutations in pre-leukemic stem cells in acute myeloid leukemia. Int J Hematol 2013; 98: 648–657.
Jan M, Snyder TM, Corces-Zimmerman MR, Vyas P, Weissman IL, Quake SR et al. Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci Transl Med 2012; 4: 149ra118.
Bejar R, Stevenson K, Abdel-Wahab O, Galili N, Nilsson B, Garcia-Manero G et al. Clinical effect of point mutations in myelodysplastic syndromes. N Engl J Med 2011; 364: 2496–2506.
Tefferi A, Pardanani A, Lim KH, Abdel-Wahab O, Lasho TL, Patel J et al. TET2 mutations and their clinical correlates in polycythemia vera, essential thrombocythemia and myelofibrosis. Leukemia 2009; 23: 905–911.
Vannucchi AM, Lasho TL, Guglielmelli P, Biamonte F, Pardanani A, Pereira A et al. Mutations and prognosis in primary myelofibrosis. Leukemia 2013; 27: 1861–1869.
Beer PA, Delhommeau F, LeCouédic JP, Dawson MA, Chen E, Bareford D et al. Two routes to leukemic transformation after a JAK2 mutation-positive myeloproliferative neoplasm. Blood 2010; 115: 2891–2900.
Schaub FX, Looser R, Li S, Hao-Shen H, Lehmann T, Tichelli A et al. Clonal analysis of TET2 and JAK2 mutations suggests that TET2 can be a late event in the progression of myeloproliferative neoplasms. Blood 2010; 115: 2003–2007.
Roche-Lestienne C, Marceau A, Labis E, Nibourel O, Coiteux V, Guilhot J et alFi-LMC group. Mutation analysis of TET2, IDH1, IDH2 and ASXL1 in chronic myeloid leukemia. Leukemia 2011; 25: 1661–1664.
Tefferi A, Lim KH, Abdel-Wahab O, Lasho TL, Patel J, Patnaik MM et al. Detection of mutant TET2 in myeloid malignancies other than myeloproliferative neoplasms: CMML, MDS, MDS/MPN and AML. Leukemia 2009; 23: 1343–1345.
Kosmider O, Gelsi-Boyer V, Cheok M, Grabar S, Della-Valle V, Picard F et al. TET2 mutation is an independent favorable prognostic factor in myelodysplastic syndromes (MDSs). Blood 2009; 114: 3285–3291.
Itzykson R, Kosmider O, Cluzeau T, Mansat-De Mas V, Dreyfus F, Beyne-Rauzy O et al. Impact of TET2 mutations on response rate to azacitidine in myelodysplastic syndromes and low blast count acute myeloid leukemias. Leukemia 2011; 25: 1147–1152.
Grossmann V, Kohlmann A, Eder C, Haferlach C, Kern W, Cross NC et al. Molecular profiling of chronic myelomonocytic leukemia reveals diverse mutations in >80% of patients with TET2 and EZH2 being of high prognostic relevance. Leukemia 2011; 25: 877–879.
Kohlmann A, Grossmann V, Klein HU, Schindela S, Weiss T, Kazak B et al. Next-generation sequencing technology reveals a characteristic pattern of molecular mutations in 72.8% of chronic myelomonocytic leukemia by detecting frequent alterations in TET2, CBL, RAS, and RUNX1. J Clin Oncol 2010; 28: 3858–3865.
Flach J, Dicker F, Schnittger S, Kohlmann A, Haferlach T, Haferlach C . Mutations of JAK2 and TET2, but not CBL are detectable in a high portion of patients with refractory anemia with ring sideroblasts and thrombocytosis. Haematologica 2010; 95: 518–519.
Nibourel O, Kosmider O, Cheok M, Boissel N, Renneville A, Philippe N et al. Incidence and prognostic value of TET2 alterations in de novo acute myeloid leukemia achieving complete remission. Blood 2010; 116: 1132–1135.
Weissmann S, Alpermann T, Grossmann V, Kowarsch A, Nadarajah N, Eder C et al. Landscape of TET2 mutations in acute myeloid leukemia. Leukemia 2012; 26: 934–942.
Konstandin N, Bultmann S, Szwagierczak A, Dufour A, Ksienzyk B, Schneider F et al. Genomic 5-hydroxymethylcytosine levels correlate with TET2 mutations and a distinct global gene expression pattern in secondary acute myeloid leukemia. Leukemia 2011; 25: 1649–1652.
Kosmider O, Delabesse E, de Mas VM, Cornillet-Lefebvre P, Blanchet O, Delmer A et al. GOELAMS Investigators. TET2 mutations in secondary acute myeloid leukemias: a French retrospective study. Haematologica 2011; 96: 1059–1063.
Chou WC, Chou SC, Liu CY, Chen CY, Hou HA, Kuo YY et al. TET2 mutation is an unfavorable prognostic factor in acute myeloid leukemia patients with intermediate-risk cytogenetics. Blood 2011; 118: 3803–3810.
Tefferi A, Levine RL, Lim KH, Abdel-Wahab O, Lasho TL, Patel J et al. Frequent TET2 mutations in systemic mastocytosis: clinical, KITD816V and FIP1L1-PDGFRA correlates. Leukemia 2009; 23: 900–904.
Soucie E, Hanssens K, Mercher T, Georgin-Lavialle S, Damaj G, Livideanu C et al. In aggressive forms of mastocytosis, TET2 loss cooperates with c-KITD816V to transform mast cells. Blood 2012; 120: 4846–4849.
Jardin F, Ruminy P, Parmentier F, Troussard X, Vaida I, Stamatoullas A et al. TET2 and TP53 mutations are frequently observed in blastic plasmacytoid dendritic cell neoplasm. Br J Haematol 2011; 153: 413–416.
Menezes J, Acquadro F, Wiseman M, Gonzalo Gómez-López G, Salgado RN, Talavera-Casañas JG et al. Exome sequencing reveals novel and recurrent mutations with clinical impact in Blastic Plasmacytoid Dendritic Cell Neoplasm. Leukemia 2013; e-pub ahead of print 27 September 2013 doi:10.1038/leu.2013.283.
Marcucci G, Maharry K, Wu YZ, Radmacher MD, Mrózek K, Margeson D et al. IDH1 and IDH2 gene mutations identify novel molecular subsets within de novo cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol 2010; 28: 2348–2355.
Tefferi A, Lasho TL, Abdel-Wahab O, Guglielmelli P, Patel J, Caramazza D et al. IDH1 and IDH2 mutation studies in 1473 patients with chronic-, fibrotic- or blast-phase essential thrombocythemia, polycythemia vera or myelofibrosis. Leukemia 2010; 24: 1302–1309.
McKenney AS, Levine RL . Isocitrate dehydrogenase mutations in leukemia. J Clin Invest 2013; 123: 3672–3677.
Losman JA, Looper RE, Koivunen P, Lee S, Schneider RK, McMahon C et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 2013; 339: 1621–1625.
Wang F, Travins J, DeLaBarre B, Penard-Lacronique V, Schalm S, Hansen E et al. Targeted inhibition of mutant IDH2 in leukemia cells induces cellular differentiation. Science 2013; 340: 622–626.
Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 2012; 483: 474–478.
Janin M, Mylonas E, Saada V, Micol JB, Renneville A, Quivoron C et al. Serum 2-HG production in IDH1 and IDH2 mutated de novo acute myeloid leukemia: A Study by the Acute Leukemia French Association Group. J Clin Oncol 2014; 32: 297–305.
Jankowska AM, Makishima H, Tiu RV, Szpurka H, Huang Y, Traina F et al. Mutational spectrum analysis of chronic myelomonocytic leukemia includes genes associated with epigenetic regulation: UTX, EZH2, and DNMT3A. Blood 2011; 118: 3932–3941.
Abdel-Wahab O, Adli M, LaFave LM, Gao J, Hricik T, Shih AH et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 2012; 22: 180–193.
Lemonnier F, Couronné L, Parrens M, Jaïs JP, Travert M, Lamant L et al. Recurrent TET2 mutations in peripheral T-cell lymphomas correlate with TFH-like features and adverse clinical parameters. Blood 2012; 120: 1466–1469.
Couronné L, Bastard C, Bernard OA . TET2 and DNMT3A mutations in human T-cell lymphoma. N Engl J Med 2012; 366: 95–96.
Cairns RA, Iqbal J, Lemonnier F, Kucuk C, de Leval L, Jais JP et al. IDH2 mutations are frequent in angioimmunoblastic T-cell lymphoma. Blood 2012; 119: 1901–1903.
Meissner B, Kridel R, Lim RS, Rogic S, Tse K, Scott DW et al. The E3 ubiquitin ligase UBR5 is recurrently mutated in mantle cell lymphoma. Blood 2013; 121: 3161–3164.
Asmar F, Punj V, Christensen J, Pedersen MT, Pedersen A, Nielsen AB et al. Genome-wide profiling identifies a DNA methylation signature that associates with TET2 mutations in diffuse large B-cell lymphoma. Haematologica 2013; e-pub ahead of print 12 July 2013.
Nickerson ML, Im KM, Misner KJ, Tan W, Lou H, Gold B et al. Somatic alterations contributing to metastasis of a castration-resistant prostate cancer. Hum Mutat 2013; 34: 1231–1241.
Yang H, Liu Y, Bai F, Zhang JY, Ma SH, Liu J et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013; 32: 663–669.
Pollyea DA, Raval A, Kusler B, Gotlib JR, Alizadeh AA, Mitchell BS . Impact of TET2 mutations on mRNA expression and clinical outcomes in MDS patients treated with DNA methyltransferase inhibitors. Hematol Oncol 2011; 29: 157–160.
Voso MT, Fabiani E, Piciocchi A, Matteucci C, Brandimarte L, Finelli C et al. Role of BCL2L10 methylation and TET2 mutations in higher risk myelodysplastic syndromes treated with 5-azacytidine. Leukemia 2011; 25: 1910–1913.
Braun T, Itzykson R, Renneville A, de Renzis B, Dreyfus F, Laribi K et al. Molecular predictors of response to decitabine in advanced chronic myelomonocytic leukemia: a phase 2 trial. Blood 2011; 118: 3824–3831.
Thol F, Friesen I, Damm F, Yun H, Weissinger EM, Krauter J et al. Prognostic significance of ASXL1 mutations in patients with myelodysplastic syndromes. J Clin Oncol 2011; 29: 2499–2506.
Zuber J, Shi J, Wang E, Rappaport AR, Herrmann H, Sison EA et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 2011; 478: 524–528.
Chen L, Deshpande AJ, Banka D, Bernt KM, Dias S, Buske C et al. Abrogation of MLL-AF10 and CALM-AF10-mediated transformation through genetic inactivation or pharmacological inhibition of the H3K79 methyltransferase Dot1l. Leukemia 2013; 27: 813–822.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 2013; 122: 3616–3627.
Acknowledgements
ES, OAB and WV are heading teams independently supported by the Ligue Nationale Contre le Cancer (Equipes labellisées) and receive grants from Inserm, Agence National de la Recherche and Institut National du Cancer. We are grateful to all members of Inserm UMR895 and UMR1009 who were involved in cited works or provided helpful comments. We apologize to all colleagues whose work was not cited because of space restrictions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Solary, E., Bernard, O., Tefferi, A. et al. The Ten-Eleven Translocation-2 (TET2) gene in hematopoiesis and hematopoietic diseases. Leukemia 28, 485–496 (2014). https://doi.org/10.1038/leu.2013.337
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2013.337
Keywords
This article is cited by
-
Lower Levels of TET2 Gene Expression, with a Higher Level of TET2 Promoter Methylation in Patients with AML; Evidence for the Role of Aberrant Methylation in AML Pathogenesis
Indian Journal of Hematology and Blood Transfusion (2024)
-
Alterations of the expression of TET2 and DNA 5-hmC predict poor prognosis in Myelodysplastic Neoplasms
BMC Cancer (2023)
-
Crosstalk between DNA methylation and hypoxia in acute myeloid leukaemia
Clinical Epigenetics (2023)
-
Nuclear localization of TET2 requires β-catenin activation and correlates with favourable prognosis in colorectal cancer
Cell Death & Disease (2023)
-
Pathological and genomic features of myeloproliferative neoplasms associated with splanchnic vein thrombosis in a single-center cohort
Annals of Hematology (2023)