Abstract
Background
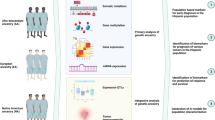
Gene polymorphisms that affect nucleotide excision repair (NER) pathway may link with higher susceptibility of breast cancer (BC); however, the significance of these associations may vary conferring to the individual ethnicity. Xeroderma pigmentosum complementation gene (XPC) plays a substantial role in recognizing damaged DNA during NER process.
Objective and methods
To estimate the relationship among XPC polymorphisms and breast cancer (BC) risk, we carried out a case–control-association study with 493 BC cases and 387 controls using TETRA–ARMS-PCR. Distributional differences of clinical features, demographic factors and XPC polymorphisms among BC cases and controls were examined by conditional logistic regression model. Kaplan–Meier test was applied to predict survival distributions and protein structure was predicted using computational tools.
Results
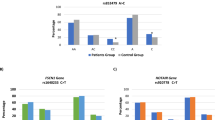
Obesity, consanguinity, positive marital status and BC family history were associated (P ≤ 0.01) with higher BC risk. Genotyping revealed significant involvement (P ≤ 0.01) of two XPC polymorphisms rs2228001–A > C (OR = 3.8; CI 1.9–7.6) and rs2733532–C > T (OR = 2.6; CI 1.4–5.03) in BC development, asserting them potential risk factors for increased BC incidence. However, no association (P > 0.05) was detected for overall or progression free survival for both XPC polymorphisms possibly due to shorter follow-up time (45 months). As compared to normal XPC structure, pronounced conformational changes have been observed in the C-terminus of XPCQ939K, bearing rs2228001–A > C substitution. In XPCQ939K, two additional α-helices were observed at A292-E297 and Y252-R286, while L623-M630 and L649-L653 helices were converted into loop conformation.
Conclusion
In conclusion, both XPC polymorphisms confer significant association with increased BC risk. rs2228001 substitution may change the structural and functional preferences of XPC C-terminus, while rs2733532 may have regulatory role thereby leading to potential BC risk.




Similar content being viewed by others
References
He B-s, Xu T, Pan Y-q, Wang H-j, Cho WC, Lin K, Sun H-l, Gao T-y, Wang S-k. Nucleotide excision repair pathway gene polymorphisms are linked to breast cancer risk in a Chinese population. Oncotarget. 2016;7(51):84872.
Grand RJA, Reynolds JJ. DNA repair and replication: mechanisms and clinical significance. Boca Raton: CRC Press; 2018.
Pongsavee M, Wisuwan K. ERCC5 rs751402 polymorphism is the risk factor for sporadic breast cancer in Thailand. Int J Mol Epidemiol Genet. 2018;9(4):27.
Winczura A, Reynolds JJ. The repair of DNA single-strand breaks and DNA adducts: mechanisms and links to human disease. DNA repair and replication: mechanisms and clinical significance, 2018.
Lehmann J. Functional relevance of spontaneous alternative splice variants of xeroderma pigmentosum genes: Prognostic marker for skin cancer risk and disease outcome?, Georg-August-Universität Göttingen, 2017.
Schäfer A. Clinical, functional, and genetic analysis of NER defective patients and characterization of five novel XPG mutations. Niedersächsische Staats-und Universitätsbibliothek Göttingen, 2012.
Latimer JJ, Johnson JM, Kelly CM, Miles TD, Beaudry-Rodgers KA, Lalanne NA, Vogel VG, Kanbour-Shakir A, Kelley JL, Johnson RR. Nucleotide excision repair deficiency is intrinsic in sporadic stage I breast cancer. Proc Natl Acad Sci. 2010;107(50):21725–30.
Cui J, Tan H, Jiang L, Yuan W, Guan Q. Association between XPC rs2228000 (C/T) polymorphism and the susceptibility of breast cancer: a Meta-analysis. J Int Oncol. 2016;43(10):752–7.
Özgöz A, Öztürk KH, Yükseltürk A, Şamlı H, Başkan Z, İçduygu FM, Bacaksız M. Genetic variations of DNA repair genes in breast cancer. Pathol Oncol Res. 2019;25(1):107–14.
Malik SS, Mubarik S, Masood N, Khadim MT. An insight into clinical outcome of XPG polymorphisms in breast cancer. Mol Biol Rep. 2018;45(6):2369–75.
Romanowicz H, Strapagiel D, Słomka M, Sobalska-Kwapis M, Kępka E, Siewierska-Górska A, Zadrożny M, Bieńkiewicz J, Smolarz B. New single nucleotide polymorphisms (SNPs) in homologous recombination repair genes detected by microarray analysis in Polish breast cancer patients. Clin Exp Med. 2017;17(4):541–6.
Aizat AAA, Nurfatimah MSS, Aminudin MM, Ankathil R. XPC Lys939Gln polymorphism, smoking and risk of sporadic colorectal cancer among Malaysians. World J Gastroenterol. 2013;19(23):3623.
DeSantis CE, Bray F, Ferlay J, Lortet-Tieulent J, Anderson BO, Jemal A. International variation in female breast cancer incidence and mortality rates. Cancer Epidemiol Biomarkers Prev. 2015;24(10):1495–506. https://doi.org/10.1158/1055-9965.EPI-15-0535.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97(16):1180–4.
Ye S, Dhillon S, Ke X, Collins AR, Day IN. An efficient procedure for genotyping single nucleotide polymorphisms. Nucleic Acids Res. 2001;29(17):e88.
Bhagwat M. Searching NCBI’s dbSNP database. Curr Protoc Bioinform. 2010;32(1):1–19.
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ. The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc. 2015;10(6):845.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2015;12(1):7.
Abraham M, van der Spoel D, Lindahl E, Hess B. The GROMACS development team GROMACS User Manual version 5.1. SoftwareX. 2016;1:19.
Wang W, Xia M, Chen J, Deng F, Yuan R, Zhang X, Shen F. Data set for phylogenetic tree and RAMPAGE Ramachandran plot analysis of SODs in Gossypium raimondii and G. arboreum. Data Brief. 2016;9:345–8.
Wallner B, Elofsson A. Can correct protein models be identified? Protein Sci. 2003;12(5):1073–86.
Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr. 2010;66(4):486–501.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem. 2004;25(13):1605–12.
Zhang ET, He Y, Grob P, Fong YW, Nogales E, Tjian R. Architecture of the human XPC DNA repair and stem cell coactivator complex. Proc Natl Acad Sci. 2015;112(48):14817–22.
Siegel RL, Miller KD. Jemal A (2018) Cancer statistics. CA Cancer J Clin. 2018;68(1):7–30. https://doi.org/10.3322/caac.21442.
Rojas K, Stuckey A. Breast cancer epidemiology and risk factors. Clin Obstet Gynecol. 2016;59(4):651–72.
Mubarik S, Malik SS, Wang Z, Li C, Fawad M, Yu C. Recent insights into breast cancer incidence trends among four Asian countries using age-period-cohort model. Cancer Manag Res. 2019;11:8145.
Anders CK, Johnson R, Litton J, Phillips M, Bleyer A. Breast cancer before age 40 years. Semin Oncol. 2009;36(3):237–49. https://doi.org/10.1053/j.seminoncol.2009.03.001.
Butt Z, Haider SF, Arif S, Khan MR, Ashfaq U, Shahbaz U, Bukhari MH. Breast cancer risk factors: a comparison between pre-menopausal and post-menopausal women. J Pak Med Assoc. 2012;62(2):120.
Surakasula A, Nagarjunapu GC, Raghavaiah KV. A comparative study of pre- and post-menopausal breast cancer: risk factors, presentation, characteristics and management. J Res Pharm Pract. 2014;3(1):12–8. https://doi.org/10.4103/2279-042X.132704.
Aldabal BK, Koura MR. Risk factors of breast cancer among the primary health-care attendees in Eastern Saudi Arabia. Int J Med Sci Public Health. 2016;5(2):276–81.
Zahmatkesh BH, Keramat A, Alavi N, Khosravi A, Chaman R. Role of menopause and early menarche in breast cancer: a meta-analysis of Iranian studies. Nurs Midwifery Stud. 2017;6(1):e37712.
Pervaiz R, Tosun Ö, Besim H, Serakinci N. Risk factor assessment for breast cancer in North Cyprus: a comprehensive case–control study of Turkish Cypriot women. Turkish J Med Sci. 2018;48(2):293–304.
Malik SS, Baig M, Khan MB, Masood N (2019) Survival analysis of breast cancer patients with different treatments: a multi-centric clinicopathological study. JPMA.
Hinyard L, Wirth LS, Clancy JM, Schwartz T. The effect of marital status on breast cancer-related outcomes in women under 65: a SEER database analysis. Breast. 2017;32:13–7.
Cadet J, Douki T. Formation of UV-induced DNA damage contributing to skin cancer development. Photochem Photobiol Sci. 2018;17(12):1816–41.
Knijnenburg TA, Wang L, Zimmermann MT, Chambwe N, Gao GF, Cherniack AD, Fan H, Shen H, Way GP, Greene CS. Genomic and molecular landscape of DNA damage repair deficiency across The Cancer Genome Atlas. Cell Rep. 2018;23(1):239–54.
Zheng W, Cong X-F, Cai W-H, Yang S, Mao C, Zou H-W. Current evidences on XPC polymorphisms and breast cancer susceptibility: a meta-analysis. Breast Cancer Res Treat. 2011;128(3):811–5.
Hernandez-Villafuerte K, Fischer A, Latimer N. Challenges and methodologies in using progression free survival as a surrogate for overall survival in oncology. Int J Technol Assess Health Care. 2018;34(3):300–16.
Yang S, Jin T, Su H-X, Zhu J-H, Wang D-W, Zhu S-J, Li S, He J, Chen Y-H. The association between NQO1 Pro187Ser polymorphism and bladder cancer susceptibility: a meta-analysis of 15 studies. PLoS ONE. 2015;10(1):e0116500.
Long X-D, Huang H-D, Huang X-Y, Yao J-G, Xia Q. XPC codon 939 polymorphism is associated with susceptibility to DNA damage induced by aflatoxin B1 exposure. Int J Clin Exp Med. 2015;8(1):1197.
Gangwar R, Mandhani A, Mittal RD. XPC gene variants: a risk factor for recurrence of urothelial bladder carcinoma in patients on BCG immunotherapy. J Cancer Res Clin Oncol. 2010;136(5):779–86.
Krzeszinski JY, Choe V, Shao J, Bao X, Cheng H, Luo S, Huo K, Rao H. XPC promotes MDM2-mediated degradation of the p53 tumor suppressor. Mol Biol Cell. 2014;25(2):213–21.
Acknowledgements
We would like to thank all the patients, their family members and colleagues for their kind help and support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Consent for publication
The experiments were undertaken with the understanding and written consent of each subject, and that the study conforms with The Code of Ethics of the World Medical Association (Declaration of Helsinki).
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Malik, S.S., Zia, A., Rashid, S. et al. XPC as breast cancer susceptibility gene: evidence from genetic profiling, statistical inferences and protein structural analysis. Breast Cancer 27, 1168–1176 (2020). https://doi.org/10.1007/s12282-020-01121-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-020-01121-z